Admin Guide: Import Data Using The Rub Importer
⚠️ Note: This page is within the system namespace/directory because as of 2026-03-31, the RUB importer in ReSeeD only supports one set of credentials for accessing S3 buckets for any data on the system as well as data to be imported (configured using the S3_ENDPOINT, S3_ACCESS_KEY, S3_SECRET_KEY, and S3_REGION variables in the .env file). We also suggest to use the S3 bucket configured in the S3_FILE_UPLOAD_BUCKET variable in the .env file for data ingest. Because access to to S3 used by the ReSeeD service requires sysadmin access anyway, this page belongs into the system directory of the wiki for the time being.
About this process
This process uses the Bulkrax CSV from S3 parser to do imports. Metadata is prepared in CSV files, data for each dataset is provided in distinct folders (see below).
Prepare the data
The data to be imported needs to have the following file structure
-
Metadata for all datasets to be imported is read from a single
metadata.csv- The format of the columns in the
metadata.csvfile is explained in The metadata CSV format section (below). ⚠️ Make sure to take into account any known issues mentioned there! - General csv remarks
- Commas are used to separate columns, and semicolons used (in some cases) to separate values within a single column. Make sure to set the field separator accordingly.
- All text containing commas or semicolon not meant to be interpreted as separators (e.g. in description or when listing contributors by "LAST_NAME, FORENAME(S)") needs to be wrapped in quotation marks.
- Encoding: UTF-8 without BOM is advised.
- The CSV file needs to contain one row for each dataset to be imported
- Each row needs to mention the path to the dataset relative to the directory containing the
metadata.csvin the columndataset_path.
- Each row needs to mention the path to the dataset relative to the directory containing the
- The format of the columns in the
-
Within each dataset path, there needs to be a directory named
datawhere all the data for the dataset to be ingested is placed.An example data structure for 2 datasets is shown below
cl-reseed_import/set1/ ├── dataset1 │ └── data │ └── 1529 │ ├── folder_1 │ │ ├── another_file.exe │ │ └── some_other_file.json │ ├── my_software.exe │ └── mydata.json ├── dataset2 │ └── data │ ├── AV02CP07GI0 │ │ ├── anat │ │ │ └── sub-AV02CP07GI0_T1w.nii │ │ └── func │ │ └── sub-AV02CP07GI0_task-rest_bold.nii │ ├── CHANGES │ ├── README │ ├── dataset_description.json │ ├── participants.json │ └── participants.tsv └── metadata.csv -
The example zip file Example_RUB_import_data.zip has the datasets and the
metadata.csvstructured as needed.
Steps to run an import
-
Upload the data you want to import (for example: the unzipped data in Example_RUB_import_data.zip) into any S3 bucket that ReSeed has access to. We advise to use the bucket configured in the
S3_FILE_UPLOAD_BUCKETvariable in the.envfile.For example:
cl-reseed_import- make sure to make exact note of this as you will need the exact bucket name to start the import. -
Log into ReSeeD as an administrator.
-
On the dashboard you should see the options Importers and Exporters. Click on Importers.
-
In the importers page, click on
Newon the top left corner. This would open the importer form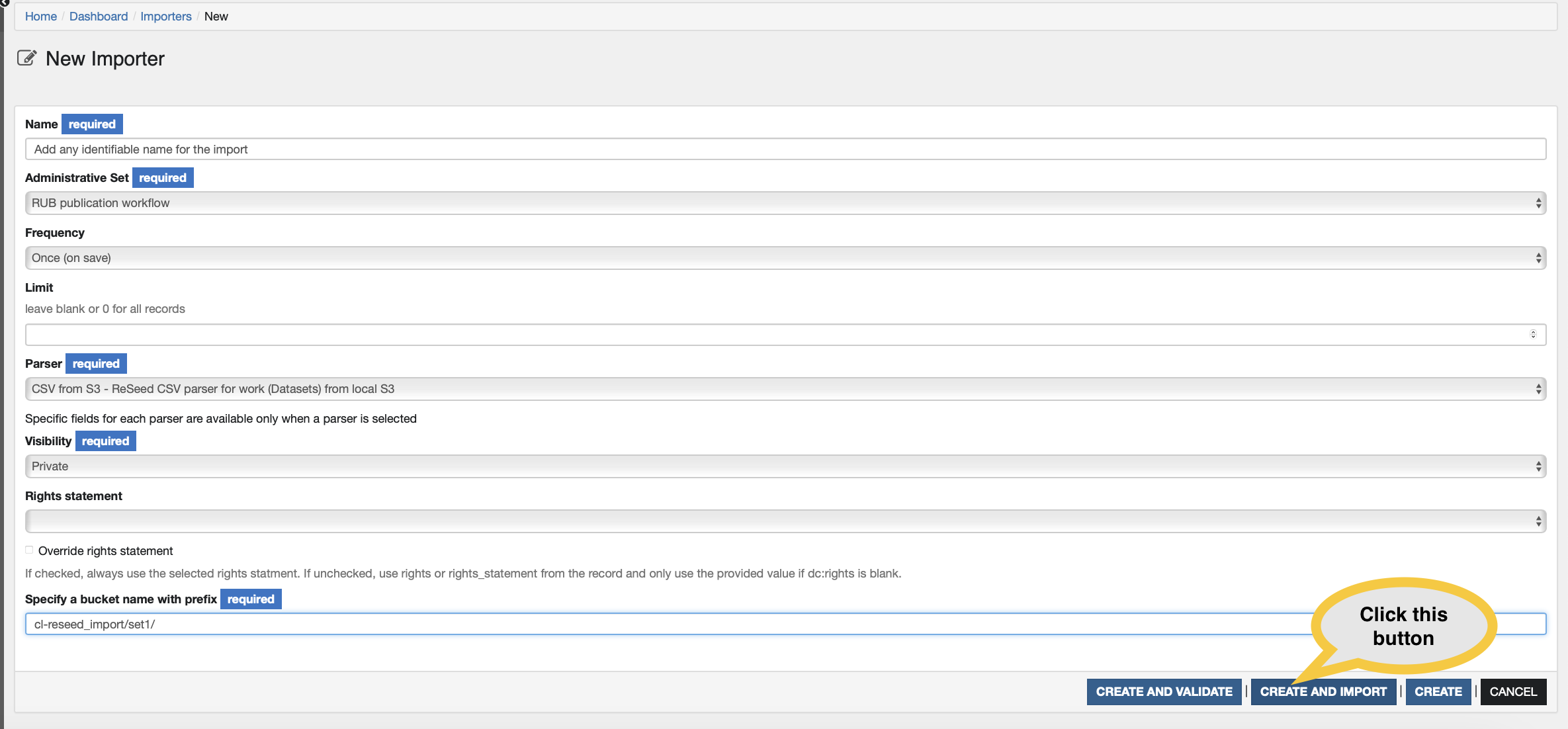
-
Fill in the Importer form with the values as indicated in the table below and click on Create and import
⚠️
CREATE AND VALIDATEandCREATEare not supported at the moment.⚠️ The RUB importer will always display
Error: NoMethodError - undefined methodcomplex_subjects'`. ⚠️ You may safely ignore this because the complex_subjects methods is only part of the CRC Importer. Look at Sidekiq to see if the importer is running succesfully.
| Field name | Value | Note |
|---|---|---|
| Name | Any identifiable name for the import unique to your system | Any identifiable name for the import |
| Administrative Set | RUB publication workflow | This will apply this workflow to all imported datasets. |
| Frequency | Once | We are running a one off import |
| Limit | 0 or leave blank | This will import all records in the metadata.csv file |
| Parser | CSV from S3 - ReSeed CSV parser for work (Datasets) from local S3 | This will choose the parser for ReSeed |
| Visibility | Private | All datasets should go through their respective worfklow to be published |
| Rights statement | Leave blank - currently not supported | It will pick up the rights statement from the csv file |
| Specify a bucket name with prefix | cl-reseed_import | The bucket name with the prefix. You could also add a path within the bucket, for example: cl-reseed_import/set1 |
The metadata CSV format
| Column header | Cardinality | Format | Example 1 | Example 2 |
|---|---|---|---|---|
| title | One | String The title of the dataset. ⚠️ If a dataset of the same new exists already, the importer will attempt overwrite data and metadata in the dataset |
Test dataset 1 for import | Test dataset 2 for import |
| dataset_path | One | String Folder path within the bucket |
dataset1 | dataset2 |
| alternative_title | Zero or more | String The alternative title(s) of the dataset. Multiple values should be separated with a semicolon. ⚠️ Currently written to single string |
The rhythms of old men who hit things with sticks | The rhythms of old men who hit things with sticks; Huh? |
| description | Zero or one | String Description of the dataset |
A collection of rhythms from veteran rock drummers | A collection of rhythms from veteran rock drummers |
| contributor | Zero or more | Names should be entered in the format: LAST_NAME, FORENAME(S) and need to be quoted (to escape commas). Multiple contributors should be separated with a semicolon. The order of names is significant in relating them to: contributor_orcid contributor_affiliation. ⚠️ There is a known issue with renaming contributors after import. We suggest adding them manually after import! ⚠️ Currently it is not possible to define contributor roles during import |
"Starr, Ringo; Bonham, John; Densmore, John; Moon, Keith" | "Starr, Ringo; Bonham, John; Densmore, John; Moon, Keith" |
| contributor_orcid | Zero or more | ORCID IDs should be entered in their full https format URI. The order of ORCID IDs is significant in relating them to contributor. ORCID IDs should be separated with a semicolon. It should ideally have the same number of semicolons as contributor. | ;;https://orcid.org/0000-0001-5109-3700; | https://orcid.org/0000-0001-0001-3700;;; |
| contributor_affiliation | Zero or more | String The order of affiliations is significant in relating them to contributor. Affiliations should be separated with a semicolon. It should ideally have the same number of semicolons as contributor. |
The Beatles; Led Zeppelin; The Doors; The Who | The Beatles;;The Doors; |
| creator | One or more | Names should be entered in the format: LAST_NAME, FORENAME(S) and need to be quoted (to escape commas). Multiple creators should be separated with a semicolon. The order of names is significant in relating them to: creator_orcid creator_affiliation. ⚠️ There is a known issue with renaming creators after import. We advise to add them manually after import! As one creator is required make sure to pay close attention to the correct spelling of the name as you will not be able to change it later |
"Lennon, John" | "Lennon, John; McCartney, Paul" |
| creator_orcid | One or more | ORCID IDs should be entered in their full https format URI. The order of ORCID IDs is significant in relating them to creator. ORCID IDs should be separated with a semicolon. It should ideally have the same number of semicolons as creator. | https://orcid.org/0000-0001-5109-3700 | https://orcid.org/0000-0001-5109-3700;https://orcid.org/0000-0001-5109-3701 |
| creator_affiliation | One or more | String The order of affiliations is significant in relating them to creator Affiliations should be separated with a semicolon. It should ideally have the same number of semicolons as creator. |
The Beatles | The Beatles;The Beatles |
| keyword | One or more | String Multiple keywords should be separated with a semicolon ⚠️ Currently written to single string |
drumming | drumming; pop stars |
| resource_type | One or more | Must be one or more of: Book BookChapter Collection ComputationalNotebook ConferencePaper DataPaper Dataset Dissertation Event Image InteractiveResource Journal JournalArticle Model OutputManagementPlan PeerReview PhysicalObject Preprint Report Service Software Sound Standard Text Workflow Other If the value is not one of the allowed values, we will set it to Dataset ⚠️ Currently only one instance supported |
Dataset | Dataset |
| license | One | Must match with an entry from /data/config/authorities/licenses.yml, e.g.: http://rightsstatements.org/vocab/InC/1.0/ https://creativecommons.org/licenses/by/4.0/ https://creativecommons.org/licenses/by-sa/4.0/ https://creativecommons.org/licenses/by-nd/4.0/ https://creativecommons.org/licenses/by-nc/4.0/ https://creativecommons.org/licenses/by-nc-nd/4.0/ https://creativecommons.org/licenses/by-nc-sa/4.0/ http://creativecommons.org/publicdomain/zero/1.0/ http://creativecommons.org/publicdomain/mark/1.0/ http://www.apache.org/licenses/LICENSE-2.0 http://www.gnu.org/licenses/gpl.html http://opensource.org/licenses/MIT If the license URI is not one of the allowed values, we will ignore it. ⚠️ Currently not supported |
http://creativecommons.org/publicdomain/mark/1.0/ | http://opensource.org/licenses/MIT |
| date | Zero or more | Dates should be entered in the format: YYYY-MM-DD <DATE-TYPE>. Multiple dates should be separated with a semicolon. Each date must have a date type which must match an entry from /data/config/authorities/dates.yml: Accepted Available Copyrighted Collected Created Deposited Published Recorded Registered Submitted Updated Archived If the date type is not one of the allowed values, ReSeeD will ignore the date and the type The dates entered here are all metadata dates. The system dates are saved in create_date, date_modified, modified_date, date_uploaded The published date if entered above will be overwritten when you go through the submission and review workflow. |
2024-05-29 Created; 2024-06-10 Published | 2024-05-29 Created; 2024-06-10 Published |
| subject | Zero or more | String Multiple subjects should be separated with a semicolon ⚠️ Currently written to single string |
drumming | Drumming; music |
| language | Zero or more | String Multiple languages should be separated with a semicolon ⚠️ Currently written to single string |
English | English |
| location | Zero or more | String Multiple locations should be separated with a semicolon ⚠️ Currently written to single string |
London | |
| software_version | Zero or more | String Multiple software versions should be separated with a semicolon ⚠️ Currently written to single string |
||
| funder_identifier | Zero or more | Identifiers should be entered as full URIs. Multiple funders Identifier should be separated with a semicolon. The order of identifiers is significant in relating them to: funder_name award_number award_uri award_title ⚠️ Currently written to single string |
http://dx.doi.org/10.13039/501100001659 | http://dx.doi.org/10.13039/501100001659;http://dx.doi.org/10.13039/50110000165999 |
| funder_name | Zero or more | Multiple funder’s name should be separated with a semicolon. It should ideally have the same number of semicolons as identifier. The order of funder name is significant in relating them to: funder_identifier award_number award_uri award_title ⚠️ Currently written to single string |
DFG | DFG;RUB |
| award_number | Zero or more | Multiple Funder's award number should be separated with a semicolon. It should ideally have the same number of semicolons as identifier. The order of award number is significant in relating them to: funder_identifier funder_name award_uri award_title ⚠️ Currently written to single string |
A0001 | A0001;W3asxa3 |
| award_uri | Zero or more | Multiple Funder's award uri should be separated with a semicolon. It should ideally have the same number of semicolons as identifier. The order of award uri is significant in relating them to: funder_identifier funder_name award_number ⚠️ Currently written to single string award_title |
||
| award_title | Zero or more | Multiple Funder's award uri should be separated with a semicolon. It should ideally have the same number of semicolons as identifier. The order of award uri is significant in relating them to: funder_identifier funder_name award_number ⚠️ Currently written to single string award_title |